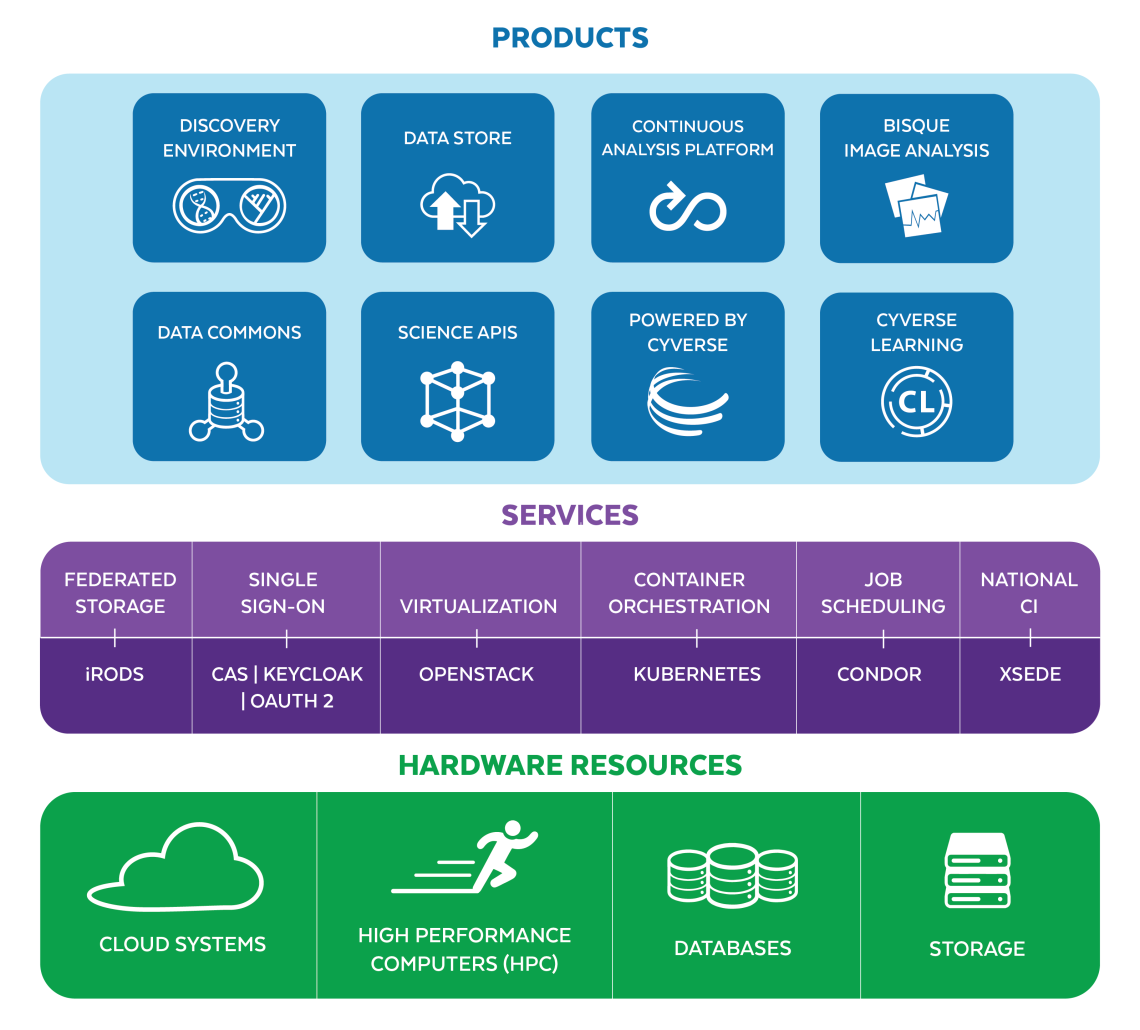
CyVerse’s powerful computational infrastructure easily handles the massive datasets and complex analyses of modern science. Offering an unprecedented degree of usability, scalability, and performance through our integrated data management system and compute resources, CyVerse provides one-stop resources for reproducible research, workflow automation, and collaboration for you, your students, and your collaborators through:

CyVerse is available anywhere with an internet connection. Whether you are a novice, educator, or experienced data scientist, our quickstarts, tutorials, webinars, workshops, and in-app chat service with our team all support your research, learning, teaching, and training needs throughout the research data lifecycle.